Software
Some listed software are open sourced. More will be open sourced as they become mature. Please check our github.com account for the latest updates.
Scientific Computing
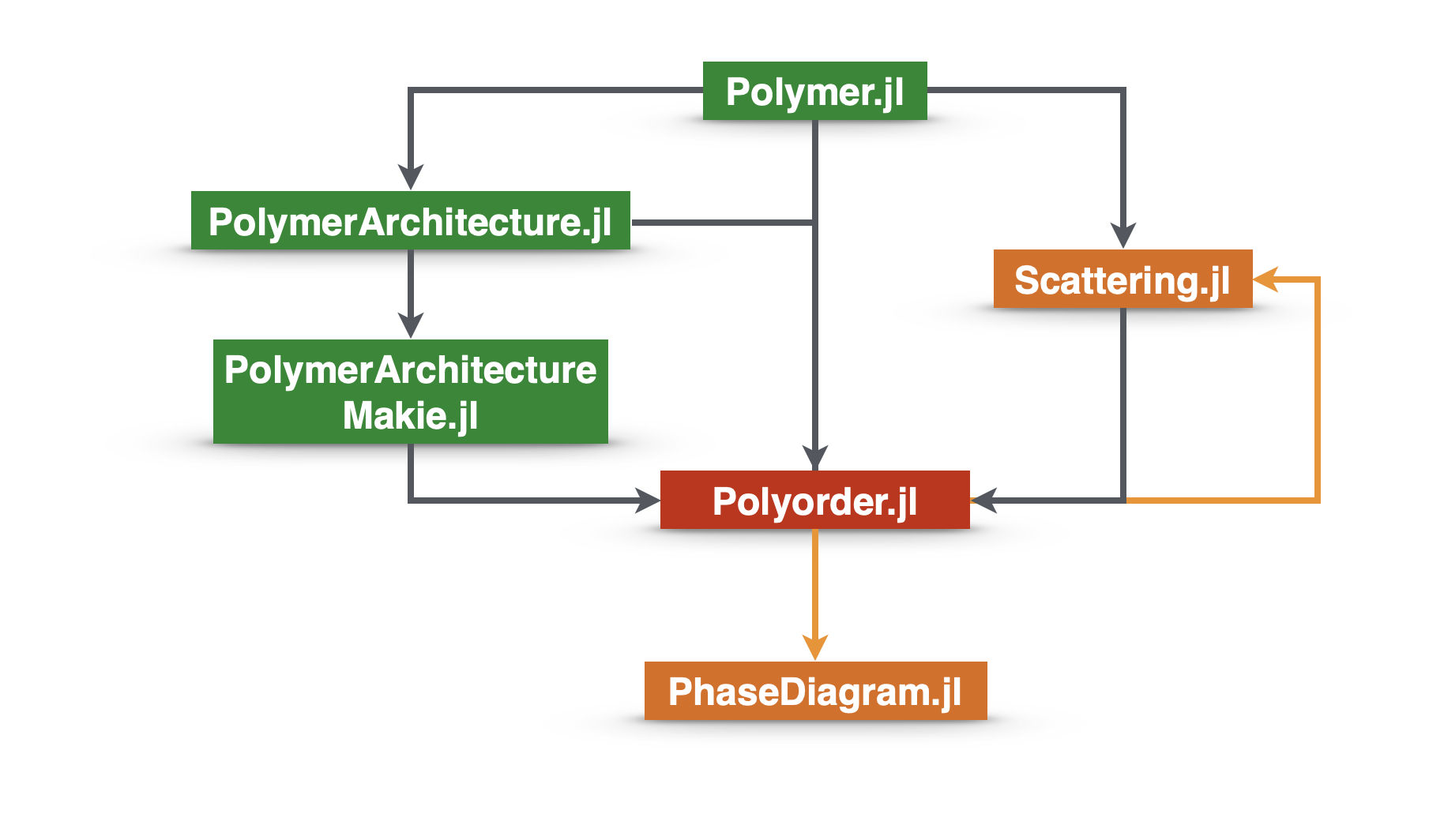
Polyorder.jl
JuliaHigh-performance, next-generation SCFT simulation platform for polymer systems
- Bleeding edge MDE solvers (ETDRK4) and preconditioned accelerators (Anderson-ETD, NGMRES, OACCEL)
- GPU acceleration: device-agnostic support via AcceleratedKernels.jl
- Low memory mode: checkpointing and shared cache for up to 80% memory reduction
- Flexible architectures for arbitrary block copolymer topologies (linear, star, comb, cyclic)
- Variable cell optimization and symmetry adapted solution for all 1D/2D/3D space groups
Polyorder Web App
JuliaWeb-based interface for running and managing Polyorder.jl simulations
- Complete web interface for SCFT simulation configuration
- Real-time simulation monitoring with live convergence plots
- Thread-safe distributed worker pool
- Block copolymer library with pre-defined architectures
- Seed library with rich set of pre-defined seeds
- User management and simulation history tracking
Scattering.jl
JuliaJulia library for computing scattering and diffraction of nanoparticles, crystals, and clusters
- Form factor of single particles and collections of particles
- Structure factor and scattering curves of crystals
- Translation, rotation, and symmetry operations on particles
- Unit cell and space group support for 1D, 2D, and 3D crystals
PhaseDiagram.jl
JuliaJulia library for computing phase diagrams of block copolymers, blends, and solutions
- Microphase separation phase boundary computation
- Macrophase separation coexistence (binodal) boundary computation
- Integration with Polyorder.jl as SCFT solver backend
Polymer.jl
JuliaCommon interface for describing polymer systems with arbitrary architectures
- Graph-based representation for arbitrary block copolymer topologies
- Convenient constructors for common polymer chains and multicomponent blends
- Serialization and configuration via YAML files
- Control parameters for systematic phase diagram construction
Installation
1
2
julia> ]
pkg> add Polymer
PolymerArchitecture.jl
JuliaGraph representation and symmetry analysis of polymer architectures
- Automatic identification of equivalent and semi-equivalent sub-architectures
- Architecture type classification (linear, star, comb, etc.)
- Symmetry reduction for polymer field-theoretic simulations (SCFT and FTS)
- Integration with Polymer.jl
CubicHermiteSpline.jl
JuliaCubic Hermite spline interpolation for 1D and 2D data
- Univariate and bivariate interpolation
- Gradient calculations
Installation
1
2
julia> ]
pkg> add CubicHermiteSpline
AcceleratedDCTs.jl
JuliaFast, device-agnostic, AbstractFFTs-compatible DCT library for 1D/2D/3D data
- High performance: optimized Makhoul's method outperforming standard separable approaches
- Device agnostic: runs on CPU (multithreaded) and GPU (CUDA, AMD via KernelAbstractions)
- VkDCT backend: VkFFT-based CUDA library for DCT-I offering ~15x speedup on GPU
- AbstractFFTs compatible with zero-allocation Plan support
- Specialized 3D kernels avoiding redundant transposes
Installation
1
2
julia> ]
pkg> add AcceleratedDCTs
CrystallographicFFT.jl
JuliaExploit crystallographic symmetry to compute FFTs on 3D periodic grids up to 28× faster
- Up to 28× speedup over full-grid FFT for highly symmetric space groups
- All 230 space groups supported
- GPU acceleration via CUDA weak extension
- R2C FFT transforms for further memory and compute savings
- SCFT-ready with built-in diffusion kernel and stress helpers
- Zero-allocation execution with pre-allocated plans
Installation
1
2
julia> ]
pkg> add CrystallographicFFT
Utility
MakiePublication.jl
JuliaPublication quality figures using Makie.jl
- Custom themes for major publishers
- 15 color palettes
- Hollow markers support
PolymerArchitectureMakie.jl
JuliaVisualization of polymer architectures using GraphMakie.jl
- Integration with PolymerArchitecture.jl
- Customizable visual styles
mpltex
PythonPython package for creating publication-quality plots
- ACS style plots
- Presentation and web styles
- Tableau color scheme
Legacy
PolyOrder
C++C++ library for polymer self-consistent field theory (deprecated)
- Non-orthogonal unit cell support
- Weakly charged polymers support
cheb++
C++C++ library for Chebyshev polynomials
- Chebyshev differential matrix
- Clenshaw-Curtis quadrature
scftpy
PythonPython package for polymer SCFT calculations
- Confined polymer simulations
- Polymer brush simulations
chebpy
PythonPython package for spectral methods of PDEs
- Chebyshev series construction
- Fast Chebyshev transform
PyDiagram
PythonPython package for generating phase diagrams
- Polyorder and PolyFTS file support
- Free energy analysis
- Phase boundary plotting
gyroid
PythonPython package for symmetry adapted basis functions
- 1D, 2D and 3D symmetry groups
- Structure renderer
NGPy
PythonWeb application for nucleation and growth simulations
- Online simulation and analysis
- Web framework for custom applications